Genetics Terms
Lets start off with a discussion about the title of my website, Rick’s SNP. A SNP (pronounced SNIP) is a Single Nucleotide Polymorphism. Nucleotides are the letters in the DNA sequence like A, C, G or T. When scientists mapped the human genome they defined what letters belonged where. A SNP happens when a persons DNA differs from the norm, or they have replaced the expected letter with a different one. This happens to everyone and on average you have about 6,000,000 of these differences between you and another person. In my case a G replaced the A (or guanine replaced adenine if you prefer) on nucleotide 845 of my HFE gene. That is my SNP. The SNP is called rs1800562(A) which is a rather clunky name. But when we look at the protein created by my SNP (genes create proteins - more on that later) the result is that the 282nd amino acid on this protein (which get named with 20 letters instead of just 4) has changed from a C to a Y (or cysteine to tyrosine). This leads to the much snappier SNP name of C282Y (also sometimes written as Cys282Tyr).
This follows for the H63D SNP which many others have. The 63rd amino acid on the HFE protein is changed from Histidine(His or H) to Aspartic Acid(Asp or D).
The 20 letters used for protein sequences each refer to a specific amino acid (which also have a 3 character abbreviation). All amino acids have the same chemical backbone and then a side chain that makes them unique. The side chain is important because the protein wants to fold into a shape. The shape is important and will affect the protein's function and viability. The shape is defined by helixes, ribbons and strings depending on what bonds that amino acids can form at their places in the sequence. The bonds are like when two magnets attract each other. The HFE gene is at the top of this page. The image can also be found here.
So sometimes a SNP can alter the shape of the protein and therefore its function. Tyrosine is a Hydrophobic amino acid which gives it specific properties, but Cysteine is special in that it has a Sulfer available at the end of its chain making it the only amino acid which can form disulfide bonds. This has a significant affect on the size, shape and function of the resulting protein. For H63D, H is a polar, uncharged molecule and D is a polar, charged. This also alters the bond that can be formed and also changes the shape and function of the protein.
We each have two copies of chromosome 6 (and therefore two copies of the HFE gene), one from each of our parents. Being Homozygous for C282Y means that both HFE genes have the same SNP on the nucleotide at location 845 while Heterozygous means that only one of the HFE genes has that SNP and the other produces a 'normal' protein.
SNPs are very common across all people's DNA. Even though it is considered a mutation this is really just the primary way that we all differ from each other. All people have SNPs. SNPs are a very common reason for the variances in peoples DNA. So SNPs can be why law enforcement agencies can tell who committed a crime based on their DNA. SNPs can affect our athletic performance, our appearance, how we respond to medications or in some cases cause disease. Generally speaking, every human has a rare SNP variant in around 150 of their 21,000 genes. Someone diagnosed with Genetic Hemochromatosis (GH) should not feel like a mutant. If having a SNP makes us a mutant then everyone is a mutant! We just happen to have a very thorough understanding of exactly what the C282Y SNP does and by the time the scientists are done we will have the same level of understanding of the other SNPs and their combinations. This is quite an undertaking and will take years to complete. Here is a good article from Nature magazine on current scientific efforts in the area of SNPs: Using SNP Data to Examine Human Phenotypic Differences
Codons are specific areas of the genetic code that are used to code amino acids. When a SNP occurs within a codon, it can alter the chemical structure of the amino acid as it does with the SNPs related to GH. The current GH related SNPs that I am aware of are available on my page discussing 23andme results - here.
Note that the current genetic tests most commonly evaluate C282Y and H63D. Some will also evaluate S65C. There will be more of these identified and their interactions are still being mapped. Again, this is why there is no negative test for GH. All of the answers are simply not yet discovered and not enough of the SNPs are being tested to say for certain that someone's Iron Homeostasis is not affected by their genetics.
23andme warning: If you appear to be a compound heterozygote (described more below), the 23andme test is not really sufficient to diagnose you as such. The reasons for this are complicated, but the basic issue is that we all have 2 of each gene and 23andme does not tell you which of those has which variant. The actual results are much different if you have all variants on one gene and none on the other, for example. So if you are a compound heterozygote, then either have your parents tested on 23andme as well or go through an additional test with a geneticist to find the full result. As it stands it is presumed that all results where there is one H63D and one C282Y variation are compound heterozygotes. This is because there are no recorded cases of a person having both H63D and C282Y variations on a single chromosome. Since it requires full exome or genome sequencing to prove this out, this assumption could prove incorrect over the course of time as more people are sequenced.
General Genetics Terms
Gene: Discrete Factors of Inheritance. Typically an area of DNA that produces one or more proteins.
Allele: Alternative Types of a Gene (A, a). These are essentially an arbitrary group of SNPs on a gene used for a specific test. Such as a c282y allele or h63d allele - without noting the other 11+ SNPs that could also exist on that gene. In purist form it would be the full DNA sequence of the gene and its promoters.
Genotype: Two alleles carried by an organism and gene
Homozygote: Two same Alleles (AA, aa)
Heterozygote: Two different Alleles (Aa)
Phenotype: Appearance, Trait
Phenotype #1 is dominant over Phenotype 2 if the F1 Hybrid has Phenotype #1 (F1 being a member of the next generation)
Autosomic recessive: Autosome means one chromosome (and not an X or Y which are allosomes), so this translates to meaning the trait is recessive when only expressed by a gene on ONE chromosome. Remember we have 23 pairs of matching chromosomes. View them all here!
Phenotypes are dominant or recessive NOT genes! Genes may control or influence many traits each of which can be dominant or recessive. So it is wrong to say that the C282Y mutation is autosomic recessive. It is correct to say that the C282Y mutation is considered autosomic recessive FOR iron overloading. It may very well be autosomic dominant for other traits like arthritis.
Wild type means what is considered normal. It is commonly abbreviated as wt or +. So research tables will frequently compare C282Y to wt or +, which just means it is comparing to those which do not have this SNP variant.
Want more? Here is a nice Genetics Glossary.
A Protein Folding Disease?
A very interesting concept for those who have followed this logic this far is that of cell stress responses. In BMC Cell Biology 2007 this article was presented by Matthew W Lawless, et al which proves that the C282Y SNP by virtue of causing a misfolded protein creates on its own a stress reaction in cells called Endoplasmic Reticulum Response as well as Unfolded Protein Response. This path of understanding may eventually explain why some symptoms of GH do not improve when iron is depleted. These symptoms may be caused by the misfolded proteins reaction in the cells themselves!
Can one parent give two copies of their HFE genes or even zero?
Genetic testing in families can have some messy results. For some people they may find that they are not genetically linked to people they think they are. This is even worse when doctors cause unnecessary confusion. This can easily happen when doctors do not provide test results correctly. For instance, I am a C282Y homozygote. My mom was tested for HFE-Hemochromatosis and the doctor told her that the result was negative. What he must have meant, of course, was that she is a C282Y heterozygote which according to the test results as they are today means that she should not be at risk for iron overload. But what she heard him say was that there is no mutation on her HFE gene. But what if she was not at least a carrier - would that mean that I must have been adopted or swapped at birth in the hospital? The answer is actually no. There is an exceptionally rare possibility of uniparental disomy and here is how it works. Chromosome passing does not always go in a textbook manner. Usually when it fails there is a severe enough issue that it causes a miscarriage. However, in the event a baby is conceived and is missing a chromosome from a parent it is possible that they can make another from the same parent. If this were to happen with chromosome 6 for instance, then one parent could pass on two copies of their chromosome 6.
Additionally, our DNA is not the same as it was when we were born! About every 5 years a new mutation enters into our bodies. Since women create all their eggs before this happens, their new mutations do not get passed along. However, men's new mutations can. Meaning that there is an even more rare possibility that if your father was tested many years before you were born and had no HFE mutation, that he could still have developed it later and passed it along to you. Much better odds of winning the lottery though.
This follows for the H63D SNP which many others have. The 63rd amino acid on the HFE protein is changed from Histidine(His or H) to Aspartic Acid(Asp or D).
The 20 letters used for protein sequences each refer to a specific amino acid (which also have a 3 character abbreviation). All amino acids have the same chemical backbone and then a side chain that makes them unique. The side chain is important because the protein wants to fold into a shape. The shape is important and will affect the protein's function and viability. The shape is defined by helixes, ribbons and strings depending on what bonds that amino acids can form at their places in the sequence. The bonds are like when two magnets attract each other. The HFE gene is at the top of this page. The image can also be found here.
So sometimes a SNP can alter the shape of the protein and therefore its function. Tyrosine is a Hydrophobic amino acid which gives it specific properties, but Cysteine is special in that it has a Sulfer available at the end of its chain making it the only amino acid which can form disulfide bonds. This has a significant affect on the size, shape and function of the resulting protein. For H63D, H is a polar, uncharged molecule and D is a polar, charged. This also alters the bond that can be formed and also changes the shape and function of the protein.
We each have two copies of chromosome 6 (and therefore two copies of the HFE gene), one from each of our parents. Being Homozygous for C282Y means that both HFE genes have the same SNP on the nucleotide at location 845 while Heterozygous means that only one of the HFE genes has that SNP and the other produces a 'normal' protein.
SNPs are very common across all people's DNA. Even though it is considered a mutation this is really just the primary way that we all differ from each other. All people have SNPs. SNPs are a very common reason for the variances in peoples DNA. So SNPs can be why law enforcement agencies can tell who committed a crime based on their DNA. SNPs can affect our athletic performance, our appearance, how we respond to medications or in some cases cause disease. Generally speaking, every human has a rare SNP variant in around 150 of their 21,000 genes. Someone diagnosed with Genetic Hemochromatosis (GH) should not feel like a mutant. If having a SNP makes us a mutant then everyone is a mutant! We just happen to have a very thorough understanding of exactly what the C282Y SNP does and by the time the scientists are done we will have the same level of understanding of the other SNPs and their combinations. This is quite an undertaking and will take years to complete. Here is a good article from Nature magazine on current scientific efforts in the area of SNPs: Using SNP Data to Examine Human Phenotypic Differences
Codons are specific areas of the genetic code that are used to code amino acids. When a SNP occurs within a codon, it can alter the chemical structure of the amino acid as it does with the SNPs related to GH. The current GH related SNPs that I am aware of are available on my page discussing 23andme results - here.
Note that the current genetic tests most commonly evaluate C282Y and H63D. Some will also evaluate S65C. There will be more of these identified and their interactions are still being mapped. Again, this is why there is no negative test for GH. All of the answers are simply not yet discovered and not enough of the SNPs are being tested to say for certain that someone's Iron Homeostasis is not affected by their genetics.
23andme warning: If you appear to be a compound heterozygote (described more below), the 23andme test is not really sufficient to diagnose you as such. The reasons for this are complicated, but the basic issue is that we all have 2 of each gene and 23andme does not tell you which of those has which variant. The actual results are much different if you have all variants on one gene and none on the other, for example. So if you are a compound heterozygote, then either have your parents tested on 23andme as well or go through an additional test with a geneticist to find the full result. As it stands it is presumed that all results where there is one H63D and one C282Y variation are compound heterozygotes. This is because there are no recorded cases of a person having both H63D and C282Y variations on a single chromosome. Since it requires full exome or genome sequencing to prove this out, this assumption could prove incorrect over the course of time as more people are sequenced.
General Genetics Terms
Gene: Discrete Factors of Inheritance. Typically an area of DNA that produces one or more proteins.
Allele: Alternative Types of a Gene (A, a). These are essentially an arbitrary group of SNPs on a gene used for a specific test. Such as a c282y allele or h63d allele - without noting the other 11+ SNPs that could also exist on that gene. In purist form it would be the full DNA sequence of the gene and its promoters.
Genotype: Two alleles carried by an organism and gene
Homozygote: Two same Alleles (AA, aa)
Heterozygote: Two different Alleles (Aa)
Phenotype: Appearance, Trait
Phenotype #1 is dominant over Phenotype 2 if the F1 Hybrid has Phenotype #1 (F1 being a member of the next generation)
Autosomic recessive: Autosome means one chromosome (and not an X or Y which are allosomes), so this translates to meaning the trait is recessive when only expressed by a gene on ONE chromosome. Remember we have 23 pairs of matching chromosomes. View them all here!
Phenotypes are dominant or recessive NOT genes! Genes may control or influence many traits each of which can be dominant or recessive. So it is wrong to say that the C282Y mutation is autosomic recessive. It is correct to say that the C282Y mutation is considered autosomic recessive FOR iron overloading. It may very well be autosomic dominant for other traits like arthritis.
Wild type means what is considered normal. It is commonly abbreviated as wt or +. So research tables will frequently compare C282Y to wt or +, which just means it is comparing to those which do not have this SNP variant.
Want more? Here is a nice Genetics Glossary.
A Protein Folding Disease?
A very interesting concept for those who have followed this logic this far is that of cell stress responses. In BMC Cell Biology 2007 this article was presented by Matthew W Lawless, et al which proves that the C282Y SNP by virtue of causing a misfolded protein creates on its own a stress reaction in cells called Endoplasmic Reticulum Response as well as Unfolded Protein Response. This path of understanding may eventually explain why some symptoms of GH do not improve when iron is depleted. These symptoms may be caused by the misfolded proteins reaction in the cells themselves!
Can one parent give two copies of their HFE genes or even zero?
Genetic testing in families can have some messy results. For some people they may find that they are not genetically linked to people they think they are. This is even worse when doctors cause unnecessary confusion. This can easily happen when doctors do not provide test results correctly. For instance, I am a C282Y homozygote. My mom was tested for HFE-Hemochromatosis and the doctor told her that the result was negative. What he must have meant, of course, was that she is a C282Y heterozygote which according to the test results as they are today means that she should not be at risk for iron overload. But what she heard him say was that there is no mutation on her HFE gene. But what if she was not at least a carrier - would that mean that I must have been adopted or swapped at birth in the hospital? The answer is actually no. There is an exceptionally rare possibility of uniparental disomy and here is how it works. Chromosome passing does not always go in a textbook manner. Usually when it fails there is a severe enough issue that it causes a miscarriage. However, in the event a baby is conceived and is missing a chromosome from a parent it is possible that they can make another from the same parent. If this were to happen with chromosome 6 for instance, then one parent could pass on two copies of their chromosome 6.
Additionally, our DNA is not the same as it was when we were born! About every 5 years a new mutation enters into our bodies. Since women create all their eggs before this happens, their new mutations do not get passed along. However, men's new mutations can. Meaning that there is an even more rare possibility that if your father was tested many years before you were born and had no HFE mutation, that he could still have developed it later and passed it along to you. Much better odds of winning the lottery though.
How HFE variants cause iron overload!
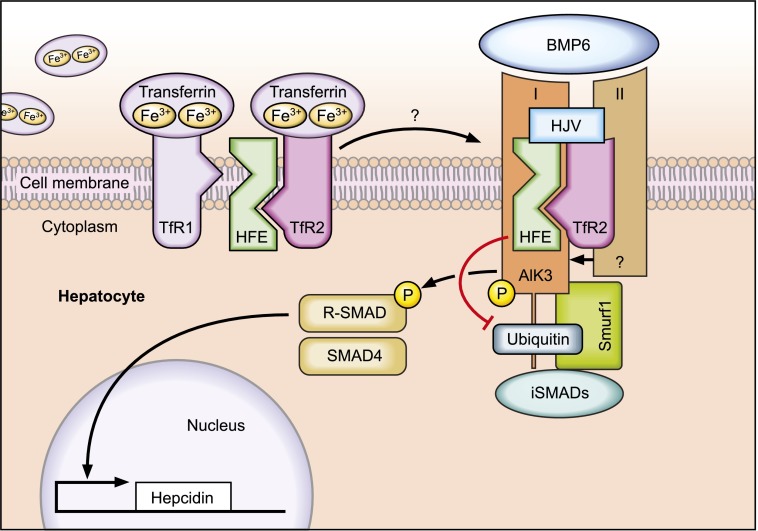
In a ground breaking paper in July of 2014 Dr Wu identified exactly how variations in the iron homeostasis process cause iron overload. The image shows HFE binding to ALK3 and expressing through the cell membrane. This must happen in order for hepcidin to be produced. In c282y, the HFE protein is not strong enough to get through the cell membrane and in h63d the amount of ALK3 available for binding actually reduces. Dr Muckenthaler explains this all better than I ever could. Her article is also the source for this image.
Here is my attempt.
Here is my attempt.
- HFE proteins are created and join with TfR1 to express through the cell membrane.
- c282y weakens HFE so it cannot get through the cell membrane.
- Less TfR1 expresses through the membrane with c282y.
- When TfR1 recognizes a high transferrin saturation % (TS%), the HFE protein breaks away from it.
- For c282y, the HFE protein acts like there is constantly a high TS%.
- The free HFE protein binds with TfR2 and stabilizes ALK3 to help it survive on the cell surface.
- Again for c282y, the HFE protein cannot get to the cell surface so ALK3 dies without it.
- For h63d (and likely s65c), HFE binds with ALK3 and they get through the cell membrane, but the altered HFE protein cannot adequately protect it and ALK3 again dies.
- Without ALK3 or with a drastically reduced amount, the process that initiates the creation of the hormone Hepcidin is either stopped or drastically reduced.
- The body does not produce an adequate amount of Hepcidin to support or reduce the existing TS%.
- The high TS% causes free iron to be deposited in places where it should not be, increases storage of iron in ferritin molecules in multiple organs and increases the amount of serum iron in the bloodstream.